Prediction of Breast Cancer Pathogenic Genes Based on Multi-agent Reinforcement Learning
-
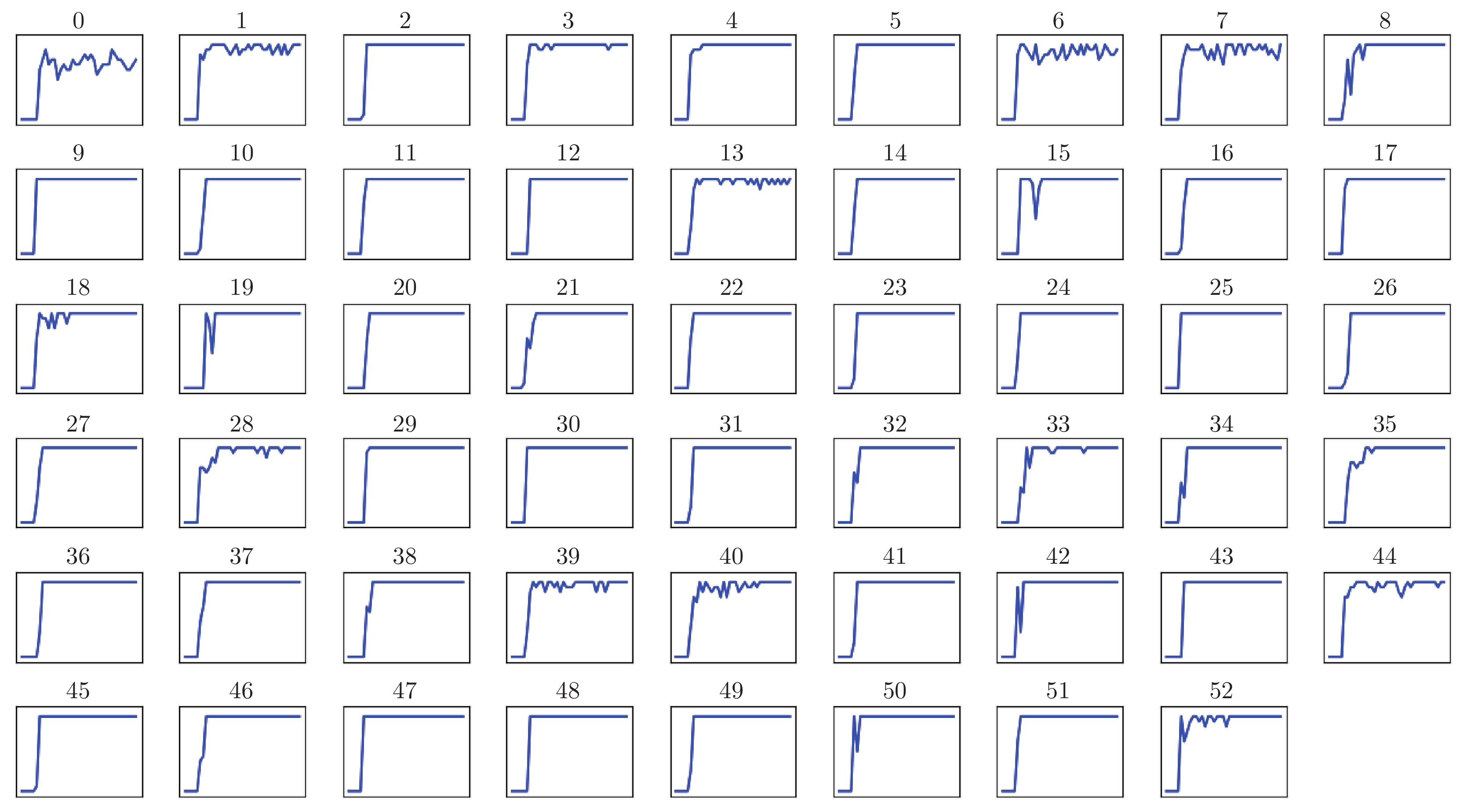
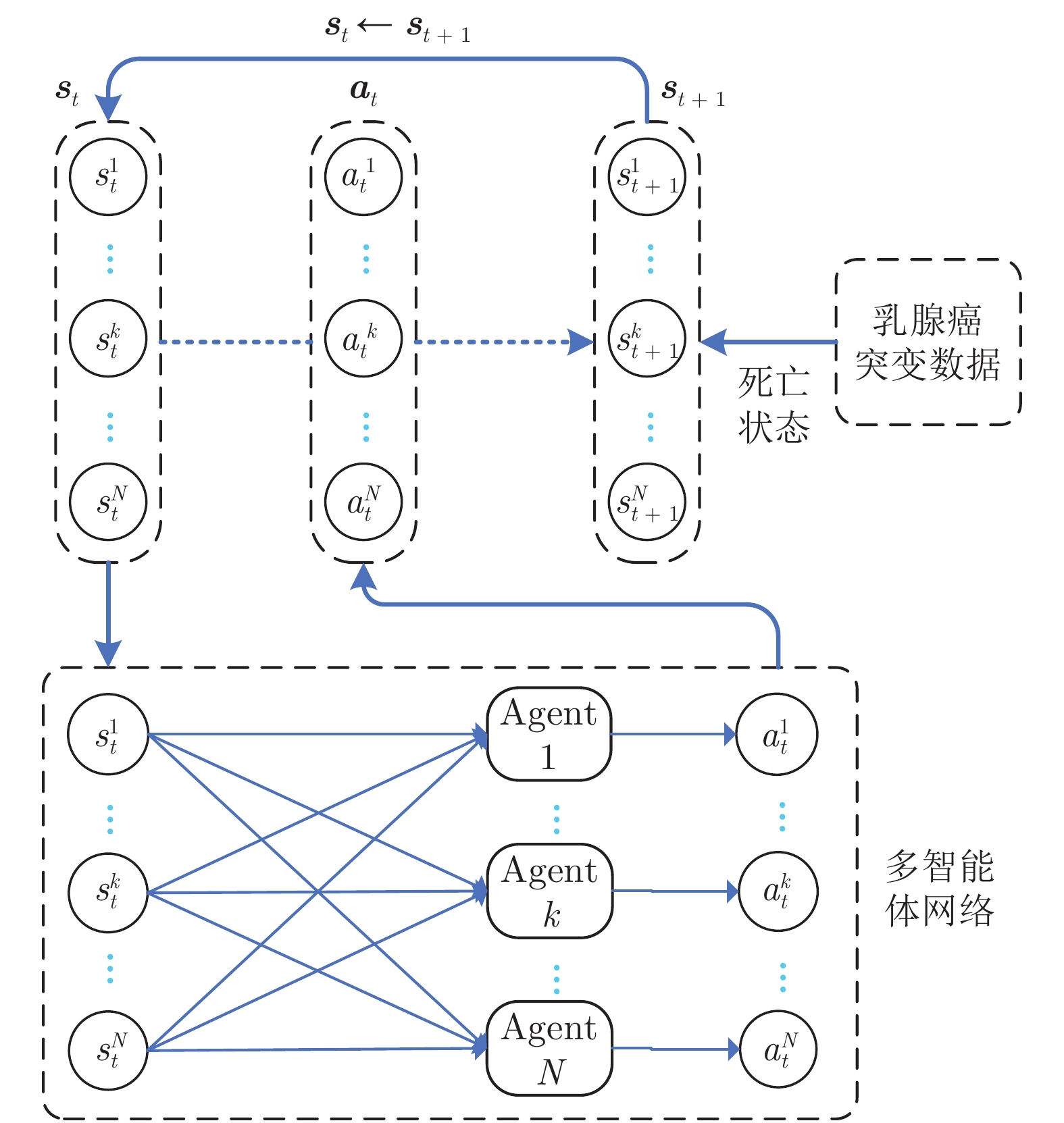
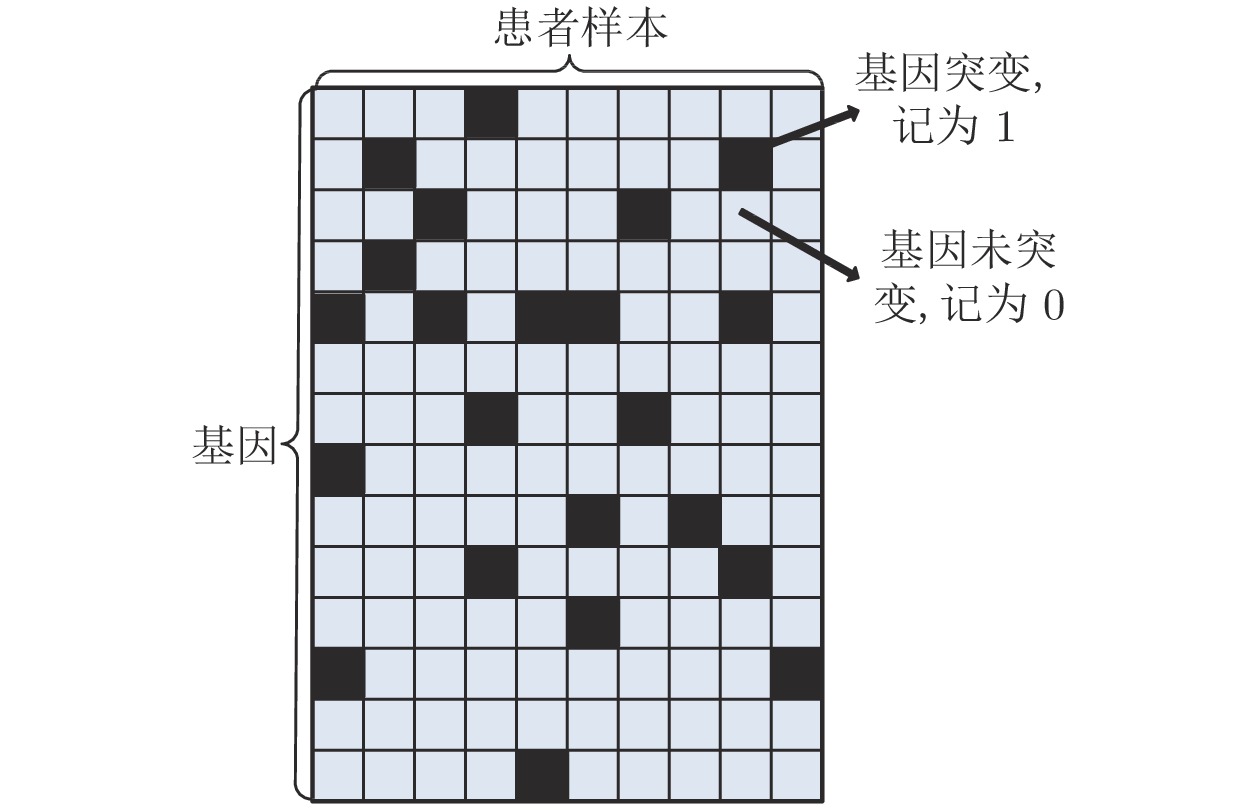
摘要: 通过分析基因突变过程, 提出利用强化学习对癌症患者由正常状态至患病状态的过程进行推断, 发现导致患者死亡的关键基因突变. 首先, 将基因视为智能体, 基于乳腺癌突变数据设计多智能体强化学习环境; 其次, 为保证智能体探索到与专家策略相同的策略和满足更多智能体快速学习, 根据演示学习理论, 分别提出两种多智能体深度Q网络: 基于行为克隆的多智能体深度Q网络和基于预训练记忆的多智能体深度Q网络; 最后, 根据训练得到的多智能体深度Q网络进行基因排序, 实现致病基因预测. 实验结果表明, 提出的多智能体强化学习方法能够挖掘出与乳腺癌发生、发展过程密切相关的致病基因.Abstract: By analyzing the gene mutation process, it is proposed to use reinforcement learning to infer the process of cancer patients from normal to disease states, and to discover the key gene mutations that lead to the death of patients. Firstly, a multi-agent reinforcement learning environment is designed based on breast cancer mutation data by viewing genes as agents. Secondly, in order to ensure that agents can find the same policy as expert policy and to satisfy more agents for rapid learning, two kinds of multi-agent deep Q networks are proposed based on demonstration learning respectively: Behavioral Cloning-based multi-agent deep Q network and pre-training memory-based multi-agent deep Q network. Finally, we sort genes according to the trained multi-agent deep Q network to achieve pathogenic gene prediction. Experimental results show that the proposed multi-agent reinforcement learning methods can dig out pathogenic genes closely related to the occurrence and development of breast cancer.
-
表 1 BCDQN和PMDQN预测的前10个致病基因
Table 1 Top 10 pathogenic genes predicted by BCDQN and PMDQN
序号 BCDQN PMDQN 1 TP53 TP53 2 FAM91A1 PIK3CA 3 TNFRSF11B TG 4 KCNQ3 HHLA1 5 MYC ASAP1 6 COL14A1 CASC8 7 CCDC26 SNORA12 8 CCN3 MYC 9 PVT1 PVT1 10 DSCC1 RN7SL329 -
[1] Wisesty U N, Mengko T R, Purwarianti A. Gene mutation detection for breast cancer disease: A review. IOP Conference Series: Materials Science and Engineering, 2020, 830(3): 032051 doi: 10.1088/1757-899X/830/3/032051 [2] Kohler S, Bauer S, Horn D, Robinson P N. Walking the interactome for prioritization of candidate pathogenic genes. The American Journal of Human Genetics, 2008, 82(4): 949-958 doi: 10.1016/j.ajhg.2008.02.013 [3] Xu B, Liu Y, Yu S, Wang L, DOng J, Lin H, et al. A network embedding model for pathogenic genes prediction by multi-path random walking on heterogeneous network. BMC Medical Genomics, 2019, 12(Suppl 10): 188 [4] Han P, Yang P, Zhao P, Shang s, Liu Y, Zhou J, et al. GCN-MF: Disease-gene association identification by graph convolutional networks and matrix factorization. In: Proceedings of the 25th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. Anchorage, AK, USA: ACM, 2019. 705−713 [5] Nagarajan N, Dhillon I S. Inductive matrix completion for predicting gene–disease associations. Bioinformatics, 2014, 30(12): i60-i68 doi: 10.1093/bioinformatics/btu269 [6] Sahu B, Mishra D. A novel feature selection algorithm using particle swarm optimization for cancer microarray data. Procedia Engineering, 2012, 38: 27-31 doi: 10.1016/j.proeng.2012.06.005 [7] Malar B, Nadarajan R, Thangam G J. A hybrid isotonic separation training algorithm with correlation-based isotonic feature selection for binary classification. Knowledge and Information Systems, 2019, 59(3): 651-683 doi: 10.1007/s10115-018-1226-6 [8] AliazKovic E, Subasi A. Breast cancer diagnosis using ga feature selection and rotation forest. Neural Computing and Applications, 2017, 28(4): 753-763 doi: 10.1007/s00521-015-2103-9 [9] Sangaiah I, Kumar A V A. Improving medical diagnosis performance using hybrid feature selection via relieff and entropy based genetic search (RF-EGA) approach: Application to breast cancer prediction. Cluster Computing, 2019, 22(3): 6899-6906 [10] Alomari O A, Khader A T, Al-Betar M A, Alyasseri Z A A. A hybrid filter-wrapper gene selection method for cancer classification. In: Proceedings of the 2018 2nd International Conference on BioSignal Analysis, Processing and Systems (ICBAPS). Porto, Portugal: IEEE, 2018. 113−118 [11] Alzubaidi A, Cosma G, Brown D, Pockley A G. Breast cancer diagnosis using a hybrid genetic algorithm for feature selection based on mutual information. In: Proceedings of 2016 International Conference on Interactive Technologies and Games (ITAG). Nottingham, UK: IEEE, 2016. 70−76 [12] Hamim M, Moudden I E, Moutachaouik H, Mustapha H. Gene selection for cancer classification: A new hybrid filter-C5.0 approach for breast cancer risk prediction. Advances in Science, Technology and Engineering Systems Journal, 2021, 6(1): 871-878 doi: 10.25046/aj060196 [13] Liu J, Su R, Zhang J, Wei L. Classification and gene selection of triple-negative breast cancer subtype embedding gene connectivity matrix in deep neural network. Briefings in Bioinformatics, DOI: 10.1093/bib/bbaa395 [14] Zhao Q, Zhang Y. Reverse construction and analysis of gene logical network based on information entropy. International Journal of Pattern Recognition and Artificial Intelligence, 2020, 34(2): 2059004 doi: 10.1142/S0218001420590041 [15] Sutton R S, Barto A G. Reinforcement learning. A Bradford Book, 1998, 15(7): 665-685 [16] 庞文砚, 范家璐, 姜艺, Lewis Frank Leroy. 基于强化学习的部分线性离散时间系统的最优输出调节. 自动化学报, DOI: 10.16383/j.aas.c190853Pang Wen-Yan, Fan Jia-Lu, Jiang Yi, Lewis Frank Leroy. Optimal output regulation of partially linear discrete-time systems using reinforcement learning. Acta Automatica Sinica10.16383/j.aas.c190853" target="_blank">, DOI: 10.16383/j.aas.c190853 [17] 施伟, 冯旸赫, 程光权, 黄红蓝, 黄金才, 刘忠, 等. 基于深度强化学习的多机协同空战方法研究. 自动化学报, 2021, 47(7): 1610−1623Shi Wei, Feng Yang-He, Cheng Guang-Quan, Huang Hong-Lan, Huang Jin-Cai, Liu Zhong, et al. Research on multi-aircraft cooperative air combat method based on deep reinforcement learning. Acta Automatica Sinica, 2021, 47(7): 1610−1623 [18] Wu T, Zhou P, Wang B, Li A, Tang X, Xu Z, et al. Joint traffic control and multi-channel reassignment for core backbone network in sdn-iot: A multi-agent deep reinforcement learning approach. IEEE Transactions on Network Science and Engineering, 2021, 8(1): 231-245 doi: 10.1109/TNSE.2020.3036456 [19] Volodymyr M, Koray K, David S, Rusu A A, Joel V, Bellemare M G, et al. Human-level control through deep reinforcement learning. Nature, 2015, 518(7540): 529-533 doi: 10.1038/nature14236 [20] Martinez D, Alenya G, Torras, C. Relational reinforcement learning with guided demonstrations. Artificial Intelligence, 2017, 247: 295-312 doi: 10.1016/j.artint.2015.02.006 [21] Torabi F, Warnell G, Stone P. Behavioral cloning from observation. In: Proceedings of the Twenty-Seventh International Joint Conference on Artificial Intelligence. Stockholm, Sweden: IJCAI, 2018. 4950−4957 [22] Silwal-Pandit L, Vollan H K M, Chin S F, Rueda O M, McKinney S, Osako T, et al. TP53 mutation spectrum in breast cancer is subtype specific and has distinct prognostic relevance. Clinical Cancer Research, 2014, 20(13): 3569-3580 doi: 10.1158/1078-0432.CCR-13-2943 [23] Funda M B, Zheng X, Shariati M, Damodaran S, Wathoo C, Brusco L, et al. Survival outcomes by TP53 mutation status in metastatic breast cancer. JCO Precision Oncology, 2018, 2: 1-15 [24] Han B, Yu J, Jia S, Liu X, Liang X, Li H. Prognostic value of the TP53 mutation location in metastatic breast cancer as detected by next-generation sequencing. Cancer Management and Research, 2021, 13: 3303-3316 doi: 10.2147/CMAR.S298729 [25] Xu J, Chen Y, Olopade O I. MYC and breast cancer. Genes and Cancer, 2010, 1(6): 629-640 doi: 10.1177/1947601910378691 [26] Terunuma A, Putluri N, Mishra P, Mathe E A, Ambs S. MYC-driven accumulation of 2-hydroxyglutarate is associated with breast cancer prognosis. Journal of Clinical Investigation, 2014, 124(1): 398-412 doi: 10.1172/JCI71180 [27] Camarda R, Zhou A Y, Kohnz R A, Balakrishnan S, Mahieu C, Anderton B, et al. Inhibition of fatty acid oxidation as a therapy for myc-overexpressing triple-negative breast cancer. Nature Medicine, 2016, 22(4): 427-432 doi: 10.1038/nm.4055 [28] Cho S W, Xu J, Sun R, Mumbach M R, Carter A C, Chen Y G, et al. Promoter of LncRNA gene PVT1 is a tumor-suppressor DNA boundary element. Cell, 2018, 173(6): 1398-1412 doi: 10.1016/j.cell.2018.03.068 [29] Tang J, Li Y, Sang Y, Yu B, Lv D, Zhang W, et al. LncRNA PVT1 regulates triple-negative breast cancer through KLF5/Beta-catenin signaling. Oncogene, 2018, 37(34): 4723-4734 doi: 10.1038/s41388-018-0310-4 [30] Wang Y, Zhou J, Wang Z, Wang P, Li S. Upregulation of SOX2 activated LncRNA PVT1 expression promotes breast cancer cell growth and invasion. Biochemical and Biophysical Research Communications, 2017, 493(1): 429-436 doi: 10.1016/j.bbrc.2017.09.005 [31] Anindya B. Abstract IA13: It takes two to tango: The PVT1-MYC alliance in human cancer. Cancer Research, 2016, 76(6): IA13 [32] Roychowdhury A, Samadder S, Das P, Mazumder D I, Chatterjee A, Addya S, et al. Deregulation of H19 is associated with cervical carcinoma. Genomics, 2019, 112(1): 961-970 [33] Li Z, Teng J, Jia Z, Zhang G, Ai X. The long non-coding RNA PCAL7 promotes prostate cancer by strengthening androgen receptor signaling. Journal of Clinical Laboratory Analysis, 2021, 35(2): e23645 -




 下载:
下载: