-
摘要: 针对特征子集区分度准则(Discernibility of feature subsets, DFS)没有考虑特征测量量纲对特征子集区分能力影响的缺陷, 引入离散系数, 提出GDFS (Generalized discernibility of feature subsets)特征子集区分度准则. 结合顺序前向、顺序后向、顺序前向浮动和顺序后向浮动4种搜索策略, 以极限学习机为分类器, 得到4种混合特征选择算法. UCI数据集与基因数据集的实验测试, 以及与DFS、Relief、DRJMIM、mRMR、LLE Score、AVC、SVM-RFE、VMInaive、AMID、AMID-DWSFS、CFR和FSSC-SD的实验比较和统计重要度检测表明: 提出的GDFS优于DFS, 能选择到分类能力更好的特征子集.Abstract: To overcome the deficiencies of the discernibility of feature subsets (DFS) which cannot take into account the influences from different attribute scales on the discernibility of a feature subset, the generalized DFS, shorted as GDFS, is proposed in this paper by introducing the coefficient of variation. The GDFS is combined with four search strategies, including sequential forward search (SFS), sequential backward search (SBS), sequential forward floating search (SFFS) and sequential backward floating search (SBFS) to develop four hybrid feature selection algorithms. The extreme learning machine (ELM) is adopted as a classification tool to guide feature selection process. We test the classification capability of the feature subsets detected by GDFS on the datasets from UCI machine learning repository and on the classic gene expression datasets, and compare the performance of the ELM classifiers based on the feature subsets by GDFS, DFS and classic feature selection algorithms including Relief, DRJMIM, mRMR, LLE Score, AVC, SVM-RFE, VMInaive, AMID, AMID-DWSFS, CFR, and FSSC-SD respectively. The statistical significance test is also conducted between GDFS, DFS, Relief, DRJMIM, mRMR, LLE Score, AVC, SVM-RFE, VMInaive, AMID, AMID-DWSFS, CFR, and FSSC-SD. Experimental results demonstrate that the proposed GDFS is superior to the original DFS. It can detect the feature subsets with much better capability in classification performance.
-
表 1 实验用UCI数据集描述
Table 1 Descriptions of datasets from UCI
数据集 样本个数 特征数 类别数 iris 150 4 3 thyroid-disease 215 5 3 glass 214 9 2 wine 178 13 3 Heart Disease 297 13 3 WDBC 569 30 2 WPBC 194 33 2 dermatology 358 34 6 ionosphere 351 34 2 Handwrite 323 256 2 表 2 GDFS+SFS与DFS+SFS算法的5-折交叉验证实验结果
Table 2 The 5-fold cross-validation experimental results of GDFS+SFS and DFS+SFS algorithms
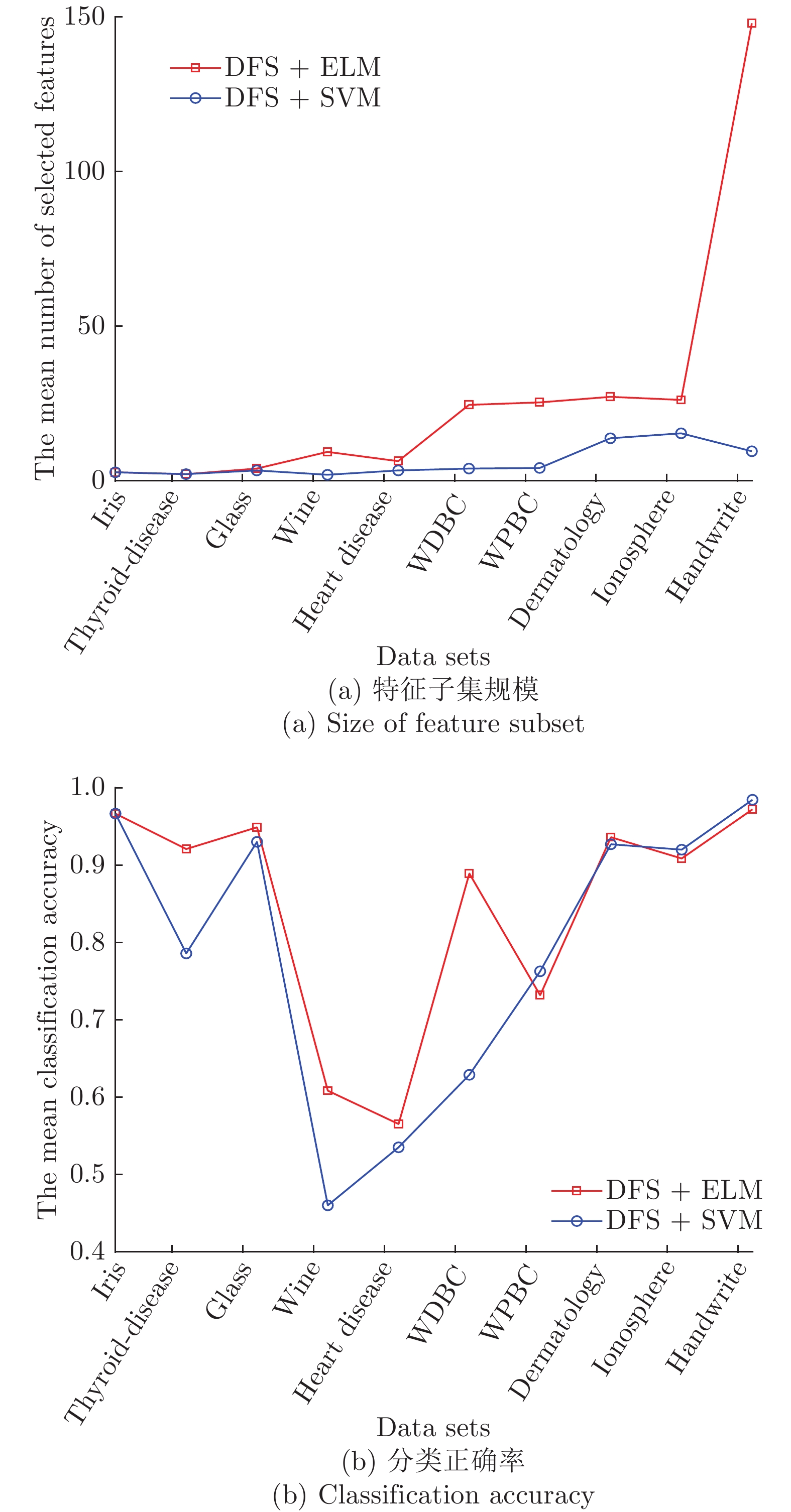
Data sets #原特征 #选择特征 测试准确率 GDFS DFS GDFS DFS iris 4 2.2 3 0.9733 0.9667 thyroid-disease 5 1.4 1.6 0.9163 0.9070 glass 9 2.4 3.2 0.9346 0.9439 wine 13 3.6 3.6 0.9272 0.8925 Heart Disease 13 2.8 3.4 0.5889 0.5654 WDBC 30 3.4 6.2 0.9227 0.9193 WPBC 33 1.8 2 0.7835 0.7732 dermatology 34 4.6 5 0.7151 0.6938 ionosphere 34 4.4 3 0.9029 0.8717 Handwrite 256 7.4 7.2 0.9657 0.9440 平均 43.1 3.4 3.82 0.8630 0.8478 表 5 GDFS+SBFS与DFS+SBFS算法的5-折交叉验证实验结果
Table 5 The 5-fold cross-validation experimental results of GDFS+SBFS and DFS+SBFS algorithms
Data sets #原特征 #选择特征 测试准确率 GDFS DFS GDFS DFS iris 4 2.4 2.8 0.98 0.9667 thyroid-disease 5 2.4 2.2 0.9395 0.9209 glass 9 5.4 4 0.8979 0.9490 wine 13 9.2 9.4 0.6519 0.6086 Heart Disease 13 5.4 6.4 0.5757 0.5655 WDBC 30 22.8 24.6 0.8911 0.8893 WPBC 33 24.6 25.4 0.7681 0.7319 dermatology 34 28.2 27.2 0.9444 0.9362 ionosphere 34 28.4 26.2 0.9174 0.9087 Handwrite 256 137.4 148 0.9938 0.9722 平均 43.1 26.62 27.62 0.8560 0.8449 表 3 GDFS+SBS与DFS+SBS算法的5-折交叉验证实验结果
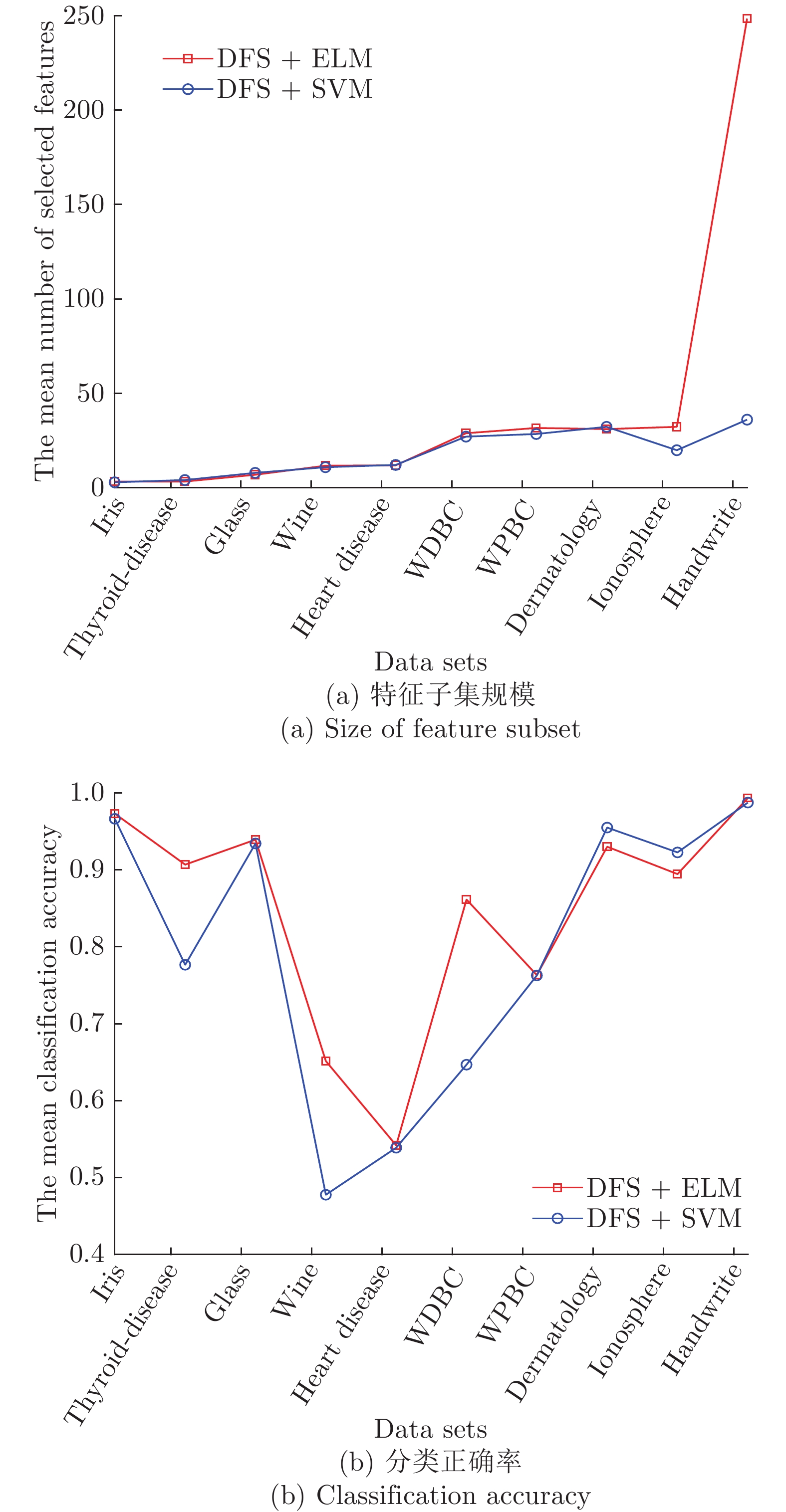
Table 3 The 5-fold cross-validation experimental results of GDFS+SBS and DFS+SBS algorithms
Data sets #原特征 #选择特征 测试准确率 GDFS DFS GDFS DFS iris 4 2.6 3.2 0.9867 0.9733 thyroid-disease 5 2.8 3.2 0.9269 0.9070 glass 9 8.2 6.8 0.9580 0.9375 wine 13 12 11.6 0.6855 0.6515 Heart Disease 13 11.8 11.8 0.5490 0.5419 WDBC 30 28 28.8 0.8981 0.8616 WPBC 33 30.8 31.6 0.7785 0.7633 dermatology 34 31 31 0.9443 0.9303 ionosphere 34 31.8 32.2 0.9031 0.8947 Handwrite 256 245 248.6 1 0.9936 平均 43.1 40.4 40.88 0.8630 0.8455 表 4 GDFS+SFFS与DFS+SFFS算法的5-折交叉验证实验结果
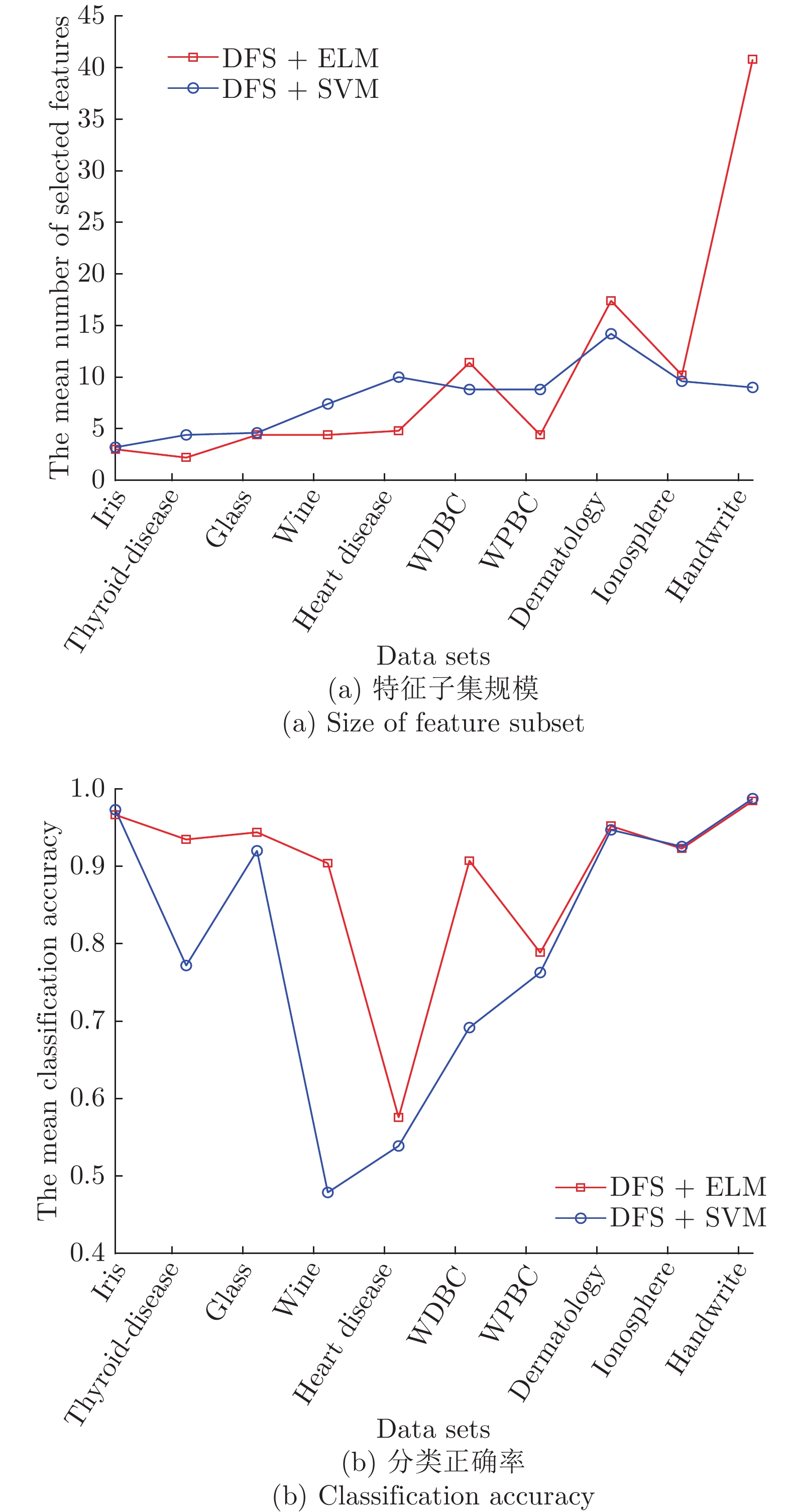
Table 4 The 5-fold cross-validation experimental results of GDFS+SFFS and DFS+SFFS algorithms
Data sets #原特征 #选择特征 测试准确率 GDFS DFS GDFS DFS iris 4 2.8 3 0.9867 0.9667 thyroid-disease 5 2.2 2.2 0.9395 0.9349 glass 9 4.2 4.4 0.9629 0.9442 wine 13 4.2 4.4 0.9261 0.9041 Heart Disease 13 4.4 4.8 0.5928 0.5757 WDBC 30 11 11.4 0.9385 0.9074 WPBC 33 5.8 4.4 0.7943 0.7886 dermatology 34 16.8 17.4 0.9522 0.9552 ionosphere 34 9.6 10.2 0.9173 0.9231 Handwrite 256 42.2 40.8 0.9907 0.9846 平均 43.1 10.32 10.3 0.8992 0.8885 表 6 实验使用的基因数据集描述
Table 6 Descriptions of gene datasets using in experiments
数据集 样本数 特征数 类别数 Colon 62 2000 2 Prostate 102 12625 2 Myeloma 173 12625 2 Gas2 124 22283 2 SRBCT 83 2308 4 Carcinoma 174 9182 11 表 7 各算法在表6基因数据集的5-折交叉验证实验结果
Table 7 The 5-fold cross-validation experimental results of all algorithms on datasets from Table 6
Data sets 算法 特征数 Accuracy AUC recall precision F-measure F2-measure Colon GDFS+SFFS 5.2 0.7590 0.8925 0.9 0.7 0.78 0.4133 DFS+SFFS 5.4 0.7256 0.78 0.8250 0.6856 0.7352 0.2332 Relief 8 0.7231 0.7575 0.9 0.6291 0.7396 0.16 DRJMIM 13 0.7282 0.7825 0.8750 0.6642 0.7495 0.3250 mRMR 5 0.7602 0.7325 0.85 0.6281 0.7185 0.1578 LLE Score 7 0.7577 0.6563 0.8750 0.6537 0.7431 0.2057 AVC 2 0.7256 0.7297 0.86 0.6439 0.7256 0.2126 SVM-RFE 5 0.7577 0.7588 0.75 0.6273 0.6775 0.3260 VMInaive 2 0.7423 1 1 0.6462 0.7848 0 AMID 8 0.7436 0.95 0.95 0.6328 0.7581 0 AMID-DWSFS 2 0.8397 0.9875 0.9750 0.6688 0.7895 0.1436 CFR 3 0.7603 0.95 1 0.6462 0.7848 0 FSSC-SD 2 0.7269 0.9750 0.9750 0.6401 0.7721 0 Prostate GDFS+SFFS 6.4 0.9305 0.9029 0.8836 0.8836 0.8829 0.8818 DFS+SFFS 6.6 0.9105 0.9349 0.8816 0.8818 0.8529 0.8497 Relief 11 0.93 0.8525 0.8255 0.7824 0.7981 0.79 DRJMIM 9 0.94 0.8629 0.7891 0.8747 0.8216 0.83 mRMR 12 0.9414 0.7895 0.7327 0.7816 0.7520 0.7597 LLE Score 26 0.9119 0.6796 0.7291 0.6582 0.6847 0.6616 AVC 12 0.9514 0.8144 0.7655 0.7598 0.7592 0.7573 SVM-RFE 22 0.92 0.8453 0.6927 0.8474 0.7567 0.7824 VMInaive 9 0.9419 0.8605 0.7655 0.7418 0.7481 0.7580 AMID 27 0.9314 0.7929 0.7655 0.7936 0.7690 0.7797 AMID-DWSFS 4 0.9514 0.7251 0.7127 0.7171 0.7011 0.7098 CFR 7 0.9410 0.7840 0.88 0.7430 0.7922 0.7942 FSSC-SD 23 0.9024 0.7796 0.8018 0.8205 0.7892 0.8130 Myeloma GDFS+SFFS 9.6 0.7974 0.6805 0.8971 0.8230 0.8558 0.5463 DFS+SFFS 9.8 0.7744 0.6296 0.8971 0.8047 0.8474 0.3121 Relief 23 0.8616 0.6453 0.8693 0.8225 0.8415 0.4631 DRJMIM 36 0.8559 0.6210 0.8392 0.7881 0.8124 0.2682 mRMR 12 0.8436 0.6332 0.8095 0.8046 0.8067 0.3539 LLE Score 64 0.8492 0.6169 0.9127 0.7909 0.8461 0.2313 AVC 22 0.8329 0.5820 0.8974 0.8098 0.8501 0.3809 SVM-RFE 20 0.8330 0.6270 0.8971 0.7935 0.8416 0.3846 VMInaive 19 0.8383 0.5639 0.8847 0.7902 0.8331 0.2691 AMID 11 0.8325 0.6743 0.8979 0.8282 0.8603 0.5254 AMID-DWSFS 38 0.8381 0.6233 0.8381 0.8197 0.8249 0.5224 CFR 14 0.8504 0.5931 0.9124 0.8014 0.8523 0.3010 FSSC-SD 15 0.8381 0.6662 0.8754 0.8173 0.8438 0.4992 Gas2 GDFS+SFFS 7.4 0.9840 0.9704 0.9051 0.9846 0.9412 0.9474 DFS+SFFS 8.4 0.9429 0.9465 0.9064 0.9212 0.9203 0.9018 Relief 4 0.9763 0.9520 0.8577 0.9316 0.8911 0.9005 DRJMIM 19 0.9750 0.9004 0.8192 0.8848 0.8449 0.8584 mRMR 5 0.9756 0.9358 0.8551 0.9131 0.8815 0.8895 LLE Score 25 0.9769 0.9312 0.8659 0.8748 0.8449 0.8538 AVC 3 0.9840 0.9073 0.8897 0.9390 0.9122 0.9160 SVM-RFE 18 0.9756 0.9009 0.8205 0.9052 0.8503 0.8716 VMInaive 10 0.9763 0.9425 0.7372 0.9778 0.8311 0.8778 AMID 16 0.9833 0.9305 0.9205 0.8829 0.8968 0.9013 AMID-DWSFS 2 0.9840 0.9247 0.8359 0.9424 0.8839 0.8977 CFR 10 0.9917 0.9080 0.9013 0.8236 0.8432 0.8434 FSSC-SD 16 0.9596 0.9095 0.8538 0.8758 0.8555 0.8642 SRBCT GDFS+SFFS 11.6 0.9372 0.9749 0.9567 0.9684 0.9579 0.9573 DFS+SFFS 11.6 0.9034 0.9130 0.9356 0.9449 0.9452 0.9352 Relief 10 0.9631 0.9479 0.9439 0.9589 0.9467 0.9390 DRJMIM 4 0.9389 0.9363 0.9656 0.9511 0.9555 0.9503 mRMR 8 0.9528 0.9479 0.9283 0.9624 0.9275 0.9294 LLE Score 11 0.9271 0.8941 0.9333 0.9332 0.9247 0.9154 AVC 8 0.9042 0.9355 0.9139 0.9544 0.9223 0.9183 SVM-RFE 13 0.8421 0.9149 0.9128 0.9385 0.9159 0.8240 VMInaive 14 0.9409 0.9181 0.9250 0.9429 0.9269 0.9188 AMID 13 0.9387 0.8999 0.9567 0.9335 0.9407 0.9239 AMID-DWSFS 9 0.9167 0.8151 0.8178 0.8516 0.82 0.7466 CFR 8 0.9314 0.6839 0.8994 0.8570 0.8693 0.7150 FSSC-SD 6 0.8806 0.9096 0.9267 0.9422 0.9284 0.9160 Carcinoma GDFS+SFFS 23.4 0.7622 0.9037 0.7872 0.7879 0.7839 0.5570 DFS+SFFS 19.4 0.7469 0.8998 0.7808 0.7869 0.7801 0.6261 Relief 42 0.7351 0.8701 0.7687 0.7785 0.7680 0.5392 DRJMIM 13 0.7757 0.8991 0.6742 0.6621 0.6656 0.4557 mRMR 24 0.8079 0.9188 0.7613 0.7505 0.7533 0.5089 LLE Score 76 0.6682 0.8452 0.6689 0.6702 0.6663 0.4109 AVC 77 0.7227 0.8746 0.7872 0.7790 0.7796 0.5068 SVM-RFE 30 0.7213 0.87 0.7027 0.6933 0.6929 0.4065 VMInaive 33 0.7443 0.8784 0.7487 0.7527 0.7441 0.4731 AMID 42 0.7307 0.8878 0.7295 0.7165 0.7194 0.4841 AMID-DWSFS 38 0.7412 0.6231 0.7558 0.7447 0.7457 0.4255 CFR 33 0.7054 0.6216 0.7514 0.74 0.7410 0.5315 FSSC-SD 21 0.7306 0.8716 0.7039 0.7016 0.6992 0.4344 表 8 各算法所选特征子集分类能力的Friedman检测结果
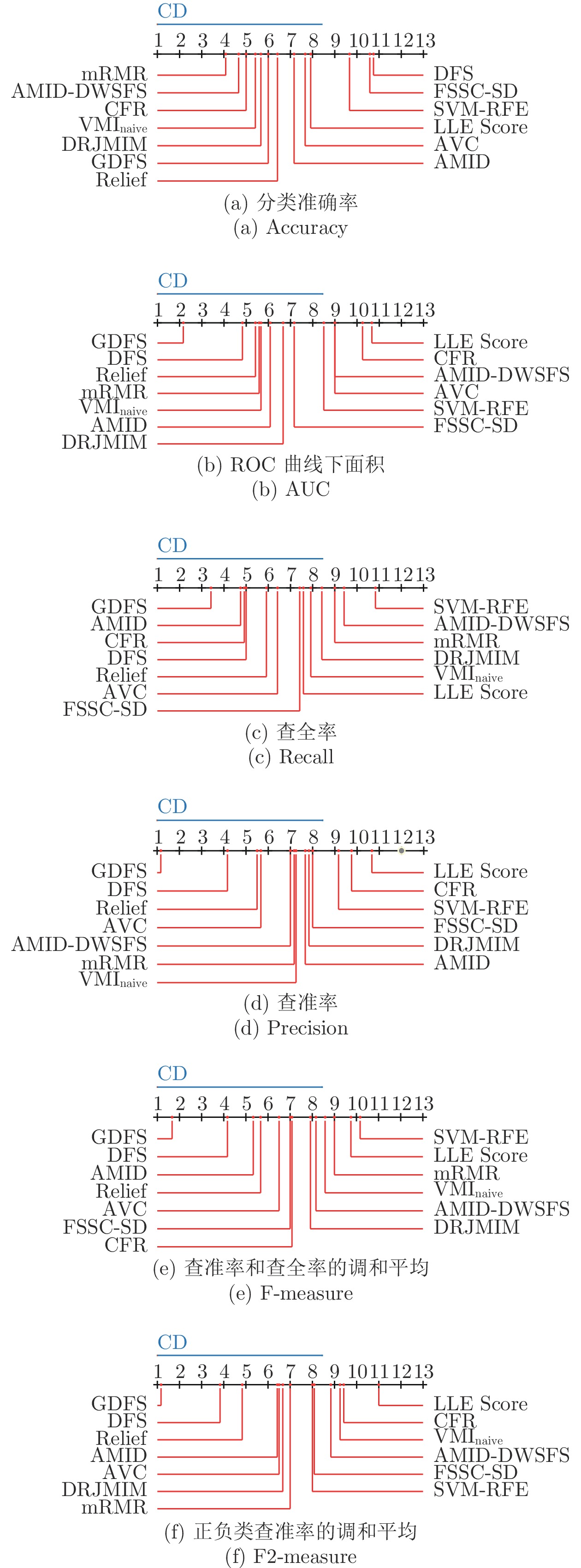
Table 8 The Friedman's test of the classification capability of feature subsets of all algorithms
Accuracy AUC recall precision F-measure F2-measure ${\chi ^2}$ 23.4094 27.5527 22.1585 29.2936 26.7608 32.5446 df 12 12 12 12 12 12 p 0.0244 0.0064 0.0358 0.0036 0.0084 0.0011 -
[1] 陈晓云, 廖梦真. 基于稀疏和近邻保持的极限学习机降维. 自动化学报, 2019, 45(2): 325-333Chen Xiao-Yun, Liao Meng-Zhen. Dimensionality reduction with extreme learning machine based on sparsity and neighborhood preserving. Acta Automatica Sinica, 2019, 45(2): 325-333 [2] Xie J Y, Lei J H, Xie W X, Shi Y, Liu X H. Two-stage hybrid feature selection algorithms for diagnosing erythemato-squamous diseases. Health Information Science and Systems, 2013, 1: Article No. 10 doi: 10.1186/2047-2501-1-10 [3] 谢娟英, 周颖. 一种新聚类评价指标. 陕西师范大学学报(自然科学版), 2015, 43(6): 1-8Xie Juan-Ying, Zhou Ying. A new criterion for clustering algorithm. Journal of Shaanxi Normal University (Natural Science Edition), 2015, 43(6): 1-8 [4] Kou G, Yang P, Peng Y, Xiao F, Chen Y, Alsaadi F E. Evaluation of feature selection methods for text classification with small datasets using multiple criteria decision-making methods. Applied Soft Computing, 2020, 86: Article No. 105836 doi: 10.1016/j.asoc.2019.105836 [5] Xue Y, Xue B, Zhang M J. Self-adaptive particle swarm optimization for large-scale feature selection in classification. ACM Transactions on Knowledge Discovery from Data, 2019, 13(5): Article No. 50 [6] Zhang Y, Gong D W, Gao X Z, Tian T, Sun X Y. Binary differential evolution with self-learning for multi-objective feature selection. Information Sciences, 2020, 507: 67-85. doi: 10.1016/j.ins.2019.08.040 [7] Nguyen B H, Xue B, Zhang M J. A survey on swarm intelligence approaches to feature selection in data mining. Swarm and Evolutionary Computation, 2020, 54: Article No. 100663 doi: 10.1016/j.swevo.2020.100663 [8] Solorio-Fernández S, Carrasco-Ochoa J A, Martínez-Trinidad J F. A review of unsupervised feature selection methods. Artificial Intelligence Review, 2020, 53(2): 907-948 doi: 10.1007/s10462-019-09682-y [9] Karasu S, Altan A, Bekiros S, Ahmad W. A new forecasting model with wrapper-based feature selection approach using multi-objective optimization technique for chaotic crude oil time series.Energy, 2020, 212: Article No. 118750 doi: 10.1016/j.energy.2020.118750 [10] Al-Tashi Q, Abdulkadir S J, Rais H, Mirjalili S, Alhussian H. Approaches to multi-objective feature selection: A systematic literature review. IEEE Access, 2020, 8: 125076-125096 doi: 10.1109/ACCESS.2020.3007291 [11] Deng X L, Li Y Q, Weng J, Zhang J L. Feature selection for text classification: A review. Multimedia Tools and Applications, 2019, 78(3): 3797-3816 doi: 10.1007/s11042-018-6083-5 [12] 贾鹤鸣, 李瑶, 孙康健. 基于遗传乌燕鸥算法的同步优化特征选择. 自动化学报, DOI: 10.16383/j.aas.c200322Jia He-Ming, Li Yao, Sun Kang-Jian. Simultaneous feature selection optimization based on hybrid sooty tern optimization algorithm and genetic algorithm. Acta Automatica Sinica, DOI: 10.16383/j.aas.c200322 [13] Xie J Y, Wang C X. Using support vector machines with a novel hybrid feature selection method for diagnosis of erythemato-squamous diseases. Expert Systems With Applications, 2011, 38(5): 5809-5815 doi: 10.1016/j.eswa.2010.10.050 [14] Bolón-Canedo V, Alonso-Betanzos A. Ensembles for feature selection: A review and future trends. Information Fusion, 2019, 52: 1-12 doi: 10.1016/j.inffus.2018.11.008 [15] Kira K, Rendell L A. The feature selection problem: Traditional methods and a new algorithm. In: Proceedings of the 10th National Conference on Artificial Intelligence. San Jos, USA: AAAI Press, 1992. 129−134 [16] Kononenko I. Estimating attributes: Analysis and extensions of RELIEF. In: Proceedings of the 7th European Conference on Machine Learning. Catania, Italy: Springer, 1994. 171−182 [17] Liu H, Setiono R. Feature selection and classification — a probabilistic wrapper approach. In: Proceedings of the 9th International Conference on Industrial and Engineering Applications of Artificial Intelligence and Expert Systems. Fukuoka, Japan: Gordon and Breach Science Publishers, 1997. 419−424 [18] Guyon I, Weston J, Barnhill S. Gene selection for cancer classification using support vector machines. Machine Learning, 2002, 46(1-3): 389-422 [19] Peng H C, Long F H, Ding C. Feature selection based on mutual information criteria of max-dependency, max-relevance, and min-redundancy. IEEE Transactions on Pattern Analysis and Machine Intelligence, 2005, 27(8): 1226-1238 doi: 10.1109/TPAMI.2005.159 [20] Chen Y W, Lin C J. Combining SVMs with various feature selection strategies. Feature Extraction: Foundations and Applications. Berlin, Heidelberg: Springer, 2006. 315−324 [21] 谢娟英, 王春霞, 蒋帅, 张琰. 基于改进的F-score与支持向量机的特征选择方法. 计算机应用, 2010, 30(4): 993-996 doi: 10.3724/SP.J.1087.2010.00993Xie Juan-Ying, Wang Chun-Xia, Jiang Shuai, Zhang Yan. Feature selection method combing improved F-score and support vector machine. Journal of Computer Applications, 2010, 30(4): 993-996 doi: 10.3724/SP.J.1087.2010.00993 [22] 谢娟英, 雷金虎, 谢维信, 高新波. 基于D-score与支持向量机的混合特征选择方法. 计算机应用, 2011, 31(12): 3292-3296Xie Juan-Ying, Lei Jin-Hu, Xie Wei-Xin, Gao Xin-Bo. Hybrid feature selection methods based on D-score and support vector machine. Journal of Computer Applications, 2011, 31(12): 3292-3296 [23] 谢娟英, 谢维信. 基于特征子集区分度与支持向量机的特征选择算法. 计算机学报, 2014, 37(8): 1704-1718Xie Juan-Ying, Xie Wei-Xin. Several feature selection algorithms based on the discernibility of a feature subset and support vector machines. Chinese Journal of Computers, 2014, 37(8): 1704-1718 [24] 李建更, 逄泽楠, 苏磊, 陈思远. 肿瘤基因选择方法LLE Score. 北京工业大学学报, 2015, 41(8): 1145-1150Li Jian-Geng, Pang Ze-Nan, Su Lei, Chen Si-Yuan. Feature selection method LLE score used for tumor gene expressive data. Journal of Beijing University of Technology, 2015, 41(8): 1145-1150 [25] Roweis S T, Saul L K. Nonlinear dimensionality reduction by locally linear embedding. Science, 2000, 290(5500): 2323-2326 doi: 10.1126/science.290.5500.2323 [26] Sun L, Wang J, Wei J M. AVC: Selecting discriminative features on basis of AUC by maximizing variable complementarity. BMC Bioinformatics, 2017, 18(Suppl 3): Article No. 50 [27] 谢娟英, 王明钊, 胡秋锋. 最大化ROC曲线下面积的不平衡基因数据集差异表达基因选择算法. 陕西师范大学学报(自然科学版), 2017, 45(1): 13-22Xie Juan-Ying, Wang Ming-Zhao, Hu Qiu-Feng. The differentially expressed gene selection algorithms for unbalanced gene datasets by maximize the area under ROC. Journal of Shaanxi Normal University (Natural Science Edition), 2017, 45(1): 13-22 [28] Hu L, Gao W F, Zhao K, Zhang P, Wang F. Feature selection considering two types of feature relevancy and feature interdependency. Expert Systems With Applications, 2018, 93: 423-434 doi: 10.1016/j.eswa.2017.10.016 [29] Sun L, Zhang X Y, Qian Y H, Xu J C, Zhang S G. Feature selection using neighborhood entropy-based uncertainty measures for gene expression data classification. Information Sciences, 2019, 502:18-41 doi: 10.1016/j.ins.2019.05.072 [30] 谢娟英, 王明钊, 周颖, 高红超, 许升全. 非平衡基因数据的差异表达基因选择算法研究. 计算机学报, 2019, 42(6): 1232-1251 doi: 10.11897/SP.J.1016.2019.01232Xie Juan-Ying, Wang Ming-Zhao, Zhou Ying, Gao Hong-Chao, Xu Sheng-Quan. Differential expression gene selection algorithms for unbalanced gene datasets. Chinese Journal of Computers, 2019, 42(6): 1232-1251 doi: 10.11897/SP.J.1016.2019.01232 [31] Li J D, Cheng K W, Wang S H, Morstatter F, Trevino R P, Tang J L, et al. Feature selection: A data perspective. ACM Computing Surveys, 2018, 50(6): Article No. 94 [32] 刘春英, 贾俊平. 统计学原理. 北京: 中国商务出版社, 2008.Liu Chun-Ying, Jia Jun-Ping. The Principles of Statistics. Beijing: China Commerce and Trade Press, 2008. [33] Huang G B, Zhu Q Y, Siew C K. Extreme learning machine: Theory and applications. Neurocomputing, 2006, 70(1-3): 489-501 doi: 10.1016/j.neucom.2005.12.126 [34] Frank A, Asuncion A. UCI machine learning repository [Online], available: http://archive.ics.uci.edu/ml, October 13, 2020 [35] Chang C C, Lin C J. LIBSVM: A library for support vector machines. ACM Transactions on Intelligent Systems and Technology, 2011, 2(3): Article No. 27 [36] Hsu C W, Chang C C, Lin C J. A practical guide to support vector classification [Online], available: https://www.ee.columbia.edu/~sfchang/course/spr/papers/svm-practical-guide.pdf, March 11, 2021 [37] Alon U, Barkai N, Notterman D A, Gish K, Ybarra S, Mack D, et al. Broad patterns of gene expression revealed by clustering analysis of tumor and normal colon tissues probed by oligonucleotide arrays. Proceedings of the National Academy of Sciences of the United States of America, 1999, 96(12): 6745-6750 doi: 10.1073/pnas.96.12.6745 [38] Singh D, Febbo P G, Ross K, Jackson D G, Manola J, Ladd C, et al. Gene expression correlates of clinical prostate cancer behavior. Cancer Cell, 2002, 1(2): 203-209 doi: 10.1016/S1535-6108(02)00030-2 [39] Tian E M, Zhan F H, Walker R, Rasmussen E, Ma Y P, Barlogie B, et al. The role of the Wnt-signaling antagonist DKK1 in the development of osteolytic lesions in multiple myeloma. The New England Journal of Medicine, 2003, 349(26): 2483-2494 doi: 10.1056/NEJMoa030847 [40] Wang G S, Hu N, Yang H H, Wang L M, Su H, Wang C Y, et al. Comparison of global gene expression of gastric cardia and noncardia cancers from a high-risk population in China. PLoS One, 2013, 8(5): Article No. e63826 doi: 10.1371/journal.pone.0063826 [41] Li W Q, Hu N, Burton V H, Yang H H, Su H, Conway C M, et al. PLCE1 mRNA and protein expression and survival of patients with esophageal squamous cell carcinoma and gastric adenocarcinoma. Cancer Epidemiology, Biomarkers & Prevention, 2014, 23(8): 1579-1588 [42] Khan J, Wei J S, Ringnér M, Saal L H, Ladanyi M, Westermann F, et al. Classification and diagnostic prediction of cancers using gene expression profiling and artificial neural networks. Nature Medicine, 2001, 7(6): 673-679 doi: 10.1038/89044 [43] Gao S Y, Steeg G V, Galstyan A. Variational information maximization for feature selection. In: Proceedings of the 30th International Conference on Neural Information Processing Systems. Barcelona, Spain: Curran Associates, 2016. 487−495 [44] Gao W F, Hu L, Zhang P, He J L. Feature selection considering the composition of feature relevancy. Pattern Recognition Letters, 2018, 112: 70-74 doi: 10.1016/j.patrec.2018.06.005 [45] 谢娟英, 丁丽娟, 王明钊. 基于谱聚类的无监督特征选择算法. 软件学报, 2020, 31(4): 1009-1024Xie Juan-Ying, Ding Li-Juan, Wang Ming-Zhao. Spectral clustering based unsupervised feature selection algorithms. Journal of Software, 2020, 31(4): 1009-1024 [46] Muschelli III J. ROC and AUC with a binary predictor: A potentially misleading metric. Journal of Classification, 2020, 37(3): 696-708 doi: 10.1007/s00357-019-09345-1 [47] Fawcett T. An introduction to ROC analysis. Pattern Recognition Letters, 2006, 27(8): 861-874 doi: 10.1016/j.patrec.2005.10.010 [48] Bowers A J, Zhou X L. Receiver operating characteristic (ROC) area under the curve (AUC): A diagnostic measure for evaluating the accuracy of predictors of education outcomes. Journal of Education for Students Placed at Risk (JESPAR), 2019, 24(1): 20-46 doi: 10.1080/10824669.2018.1523734 [49] 卢绍文, 温乙鑫. 基于图像与电流特征的电熔镁炉欠烧工况半监督分类方法. 自动化学报, 2021, 47(4): 891-902Lu Shso-Wen, Wen Yi-Xin. Semi-supervised classification of semi-molten working condition of fused magnesium furnace based on image and current features. Acta Automatica Sinica, 2021, 47(4): 891-902 [50] Xie J Y, Gao H C, Xie W X, Liu X H, Grant P W. Robust clustering by detecting density peaks and assigning points based on fuzzy weighted K-nearest neighbors. Information Sciences, 2016, 354: 19-40 doi: 10.1016/j.ins.2016.03.011 [51] 谢娟英, 吴肇中, 郑清泉. 基于信息增益与皮尔森相关系数的2D自适应特征选择算法. 陕西师范大学学报(自然科学版), 2020, 48(6): 69-81Xie Juan-Ying, Wu Zhao-Zhong, Zheng Qing-Quan. An adaptive 2D feature selection algorithm based on information gain and pearson correlation coefficient. Shaanxi Normal University (Natural Science Edition), 2020, 48(6): 69-81 -





 下载:
下载: