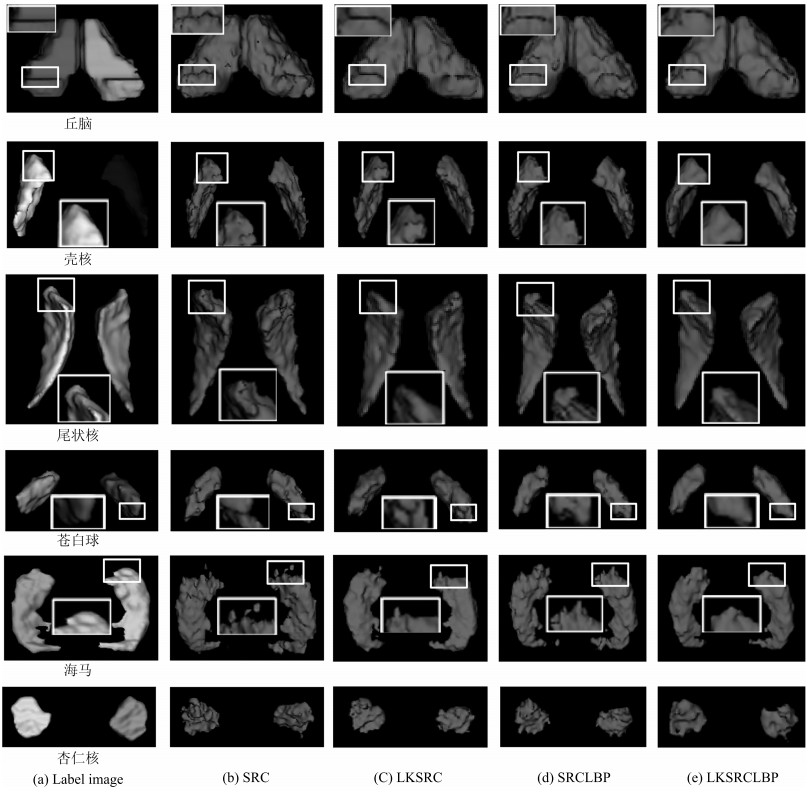
-
摘要: 深层脑结构的形态变化和神经退行性疾病相关, 对脑MR图像中的深层脑结构分割有助于分析各结构的形态变化.多图谱融合方法利用图谱图像中的先验信息, 为脑结构分割提供了一种有效的方法.大部分现有多图谱融合方法仅以灰度值作为特征, 然而深层脑结构灰度分布之间重叠的部分较多, 且边缘不明显.为克服上述问题, 本文提出一种基于线性化核多图谱融合的脑MR图像分割方法.首先, 结合纹理与灰度双重特征, 形成增强特征用于更好地表达脑结构信息.其次, 引入核方法, 通过高维映射捕获原始空间中特征的非线性结构, 增强数据间的判别性和线性相似性.最后, 利用Nyström方法, 对高维核矩阵进行估计, 通过特征值分解计算虚样本, 并在核标签融合过程中利用虚样本替代高维样本, 大大降低了核标签融合的计算复杂度.在三个公开数据集上的实验结果表明, 本文方法在较少的时间消耗内, 提高了分割精度.Abstract: Morphological changes in subcortical brain structures are related to different neurodegenerative disorders. Therefore, subcortical brain segmentation in MRI contribute to analyses of morphological changes in various structures. Multi atlas-based method provides an effective way for subcortical brain segmentation by using prior information in atlas. Most of the existing multi atlas-based methods only use intensities as features, while the distribution of gray value overlaps more and the edges of structures are not obvious. In order to solve above problems, a brain magnetic resonance image (MRI) segmentation method based on linearized kernel-based label fusion method is proposed in this paper. First, augmented features are formed by concatenating texture features and intensity features to obtain better representation. Then, kernel method is introduced to capture the nonlinear structure of features in the original space and enhance discriminability and linear similarity between features using high dimensional mapping. Finally, the Nyström method is used to estimate high dimensional kernel matrices and virtual samples are calculated by eigenvalue decomposition. Mapped samples are replaced by virtual samples in label fusion methods, which can greatly reduce the computational complexity of kernel-based label fusion methods. Experimental results on three public datasets show that the proposed method improves the segmentation accuracy with less time consumption.
-
Key words:
- Subcortical brain segmentation /
- kernel-based label fusion method /
- augmented feature /
- Nyström method /
- virtual sample
1) 本文责任编委 张道强 -
表 1 数据集基本信息
Table 1 Information of each dataset
数据集 个体数 年龄 类别数 尺寸 分辨率(mm) IBSR 18 7$\sim$71 32 $256 \times 256 \times 128$ $0.94 \times 0.94 \times 1.5$和$0.84 \times 0.84 \times 1.5$ Hammers67n 20 20$\sim$54 67 $192 \times 256 \times 124$ $0.937 \times 0.937 \times 1.5$ SATA 35 - 14 $256 \times 256 \times 287$ $1.0 \times 1.0 \times 1.0$ 表 2 LK方法在IBSR上的Dice值, 加粗的Dice值为同种LF + AF方法中的最高值
Table 2 Dice value of LK method on IBSR. Blue bolds show the maximum in the same LF + AF method
方法 丘脑 壳核 尾状核 苍白球 海马 杏仁核 平均 PB + AF 无LK 0.885 0.802 0.807 0.746 0.811 0.688 0.790 有LK ${\bf{0.899}}$ ${\bf{0.881}}$ ${\bf{0.886}}$ ${\bf{0.837}}$ ${\bf{0.816}}$ ${\bf{0.743}}$ ${\bf{0.844}}$ SRC + AF 无LK 0.864 0.823 0.809 0.758 0.714 0.639 0.768 有LK ${\bf{0.898}}$ ${\bf{0.883}}$ ${\bf{0.866}}$ ${\bf{0.831}}$ ${\bf{0.818}}$ ${\bf{0.678}}$ ${\bf{0.829}}$ DDL + AF 无LK 0.902 0.892 0.882 0.851 0.830 ${\bf{0.695}}$ 0.842 有LK ${\bf{0.905}}$ ${\bf{0.898}}$ ${\bf{0.883}}$ ${\bf{0.854}}$ ${\bf{0.839}}$ 0.678 ${\bf{0.843}}$ 表 3 LK方法在Hammers67n20上的Dice值, 加粗的Dice值为每种LF + AF方法中的最高值
Table 3 Dice value of LK method on Hammers67n20. Blue bolds show the maximum in the same LF + AF method
方法 丘脑 壳核 尾状核 苍白球 海马 杏仁核 平均 PB + AF 无LK 0.844 0.849 0.838 0.762 0.799 0.825 0.820 有LK ${\bf{0.879}}$ ${\bf{0.890}}$ ${\bf{0.889}}$ ${\bf{0.811}}$ ${\bf{0.831}}$ ${\bf{0.868}}$ ${\bf{0.861}}$ SRC + AF 无LK 0.862 0.859 0.852 0.748 0.793 0.802 0.819 有LK ${\bf{0.887}}$ ${\bf{0.885}}$ ${\bf{0.893}}$ ${\bf{0.792}}$ ${\bf{0.842}}$ ${\bf{0.839}}$ ${\bf{0.856}}$ DDL + AF 无LK 0.875 0.882 0.870 0.789 0.812 ${\bf{0.860}}$ 0.848 有LK ${\bf{0.892}}$ ${\bf{0.891}}$ ${\bf{0.875}}$ ${\bf{0.802}}$ ${\bf{0.830}}$ 0.857 ${\bf{0.858}}$ 表 4 本文方法和现有脑结构分割方法在IBSR数据集上的结果, 评价指标Dice
Table 4 Results compared with existing brain structure segmentation method, measured with Dice
FIRST FreeSurfer MS-CNN BrainSegNet FCNN M-net Dolz 本文方法 丘脑 0.889 0.840 0.889 0.89 0.87 0.90 0.92 0.905 壳核 0.875 0.809 0.875 0.91 0.83 0.90 0.90 0.898 尾状核 0.827 0.803 0.849 0.87 0.78 0.87 0.91 0.886 苍白球 0.810 0.703 0.787 0.82 0.75 0.82 0.83 0.854 海马 0.811 0.764 0.788 0.82 - 0.82 - 0.839 杏仁核 0.750 0.589 0.654 0.74 - 0.73 - 0.743 平均 0.827 0.751 0.807 0.842 0.808 0.84 0.89 0.844 -
[1] Goodro M, Sameti M, Patenaude B, Fein G. Age effect on subcortical structures in healthy adults. Psychiatry Research: Neuroimaging, 2012, 203(1): 38-45 doi: 10.1016/j.pscychresns.2011.09.014 [2] Kim M J, Hamilton J P, Gotlib I H. Reduced caudate gray matter volume in women with major depressive disorder. Psychiatry Research: Neuroimaging, 2008, 164(2): 114-122 doi: 10.1016/j.pscychresns.2007.12.020 [3] Rangini M, Jiji G W. Detection of Alzheimer$'$s disease through automated hippocampal segmentation. In: Proceedings of the 2013 Automation, Computing, Communication, Control and Compressed Sensing. Kottayam, India: IEEE, 2013. 144-149 [4] Fischl B. FreeSurfer. Neuroimage, 2012, 62(2): 774-781 doi: 10.1016/j.neuroimage.2012.01.021 [5] Patenaude B, Smith S M, Kennedy D N, Jenkinson M. A Bayesian model of shape and appearance for subcortical brain segmentation. Neuroimage, 2011, 56(3): 907-922 doi: 10.1016/j.neuroimage.2011.02.046 [6] Bao S Q, Chung A C S. Multi-scale structured CNN with label consistency for brain MR image segmentation. Computer Methods in Biomechanics and Biomedical Engineering: Imaging and Visualization, 2018, 6(1): 113-117 doi: 10.1080/21681163.2016.1182072 [7] Mehta R, Majumdar A, Sivaswamy J. BrainSegNet: a convolutional neural network architecture for automated segmentation of human brain structures. Journal of Medical Imaging, 2017, 4(2): 024003 doi: 10.1117/1.JMI.4.2.024003 [8] Shakeri M, Tsogkas S, Ferrante E. Sub-cortical brain structure segmentation using F-CNN$'$s. In: Processings of the 13th International Symposium on Biomedical Imaging. Prague, Czech Republic: IEEE, 2016. 269-272 [9] Mehta R, Sivaswamy J. M-net: a convolutional neural network for deep brain structure segmentation. In: Processings of the 14th International Symposium on Biomedical Imaging. Melbourne, Australia: IEEE, 2017. 437-440 [10] Dolz J, Desrosiers C, Ayed I B. 3D fully convolutional networks for subcortical segmentation in MRI: a large-scale study. NeuroImage, 2017, 170(2018): 456-470 http://europepmc.org/abstract/MED/28450139 [11] Shen D G, Moffat S, Resnick S M, Davatzikos C. Measuring size and shape of the hippocampus in MR images using a deformable shape model. NeuroImage, 2002, 15(2): 422-434 doi: 10.1006/nimg.2001.0987 [12] Wu G R, Kim M, Sanroma G, Wang Q, Munsell B C, Shen D G. Hierarchical multi-atlas label fusion with multi-scale feature representation and label-specific patch partition. NeuroImage, 2017, 170(2018): 456-470 [13] 田娟秀, 刘国才, 谷珊珊, 鞠忠建, 刘劲光, 顾冬冬.医学图像分析深度学习方法研究与挑战.自动化学报, 2018, 44(3): 401-424 doi: 10.16383/j.aas.2018.c170153Tian Juan-Xiu, Liu Guo-Cai, Gu Shan-Shan, Jv Zhong-Jian, Liu Jin-Guang, Gu Dong-Dong. Deep learning in medical image analysis and its challenges. Acta Automatica Sinica, 2018, 44(3): 401-424 doi: 10.16383/j.aas.2018.c170153 [14] Babalola K O, Patenaude B, Aljabar P, Schnabel J, Kennedy D, Crum W. An evaluation of four automatic methods of segmenting the subcortical structures in the brain. Neuroimage, 2009, 47(4): 1435-1447 doi: 10.1016/j.neuroimage.2009.05.029 [15] Gonzámlez-Villàm S, Oliver A, Valverde S, Wang L P, Zwiggelaar R, Lladó X. A review on brain structures segmentation in magnetic resonance imaging. Artificial Intelligence in Medicine, 2016, 73(2016): 45-69 http://www.ncbi.nlm.nih.gov/pubmed/27926381/ [16] Coupé P, Manjón J V, Fonov V, Pruessner J, Robles M, Collins D L. Patch-based segmentation using expert priors: application to hippocampus and ventricle segmentation. Neuroimage, 2011, 54(2): 940-954 doi: 10.1016/j.neuroimage.2010.09.018 [17] Rousseau F, Habas P A, Studholme C. A supervised patch-based approach for human brain labeling. IEEE Transactions on Medical Imaging, 2011, 30(10): 1852-1862 doi: 10.1109/TMI.2011.2156806 [18] Eskildsen S F, Coupé P, Fonov V, Manjó J V, Leung K K, Guizard N. BEaST: brain extraction based on nonlocal segmentation technique. Neuroimage, 2012, 59(3): 2362-2373 doi: 10.1016/j.neuroimage.2011.09.012 [19] Golts A, Elad M. Linearized kernel dictionary learning. IEEE Journal of Selected Topics in Signal Processing, 2016, 10(4): 726-739. doi: 10.1109/JSTSP.2016.2555241 [20] Li Y H, Jia F C, Qin J. Brain tumor segmentation from multimodal magnetic resonance images via sparse representation. Artificial Intelligence in Medicine, 2016, 73(2016): 1-13 http://www.ncbi.nlm.nih.gov/pubmed/27926377 [21] Tong T, Wolz R, Coupé P, Hajnal J V, Rueckert D. Segmentation of MR images via discriminative dictionary learning and sparse coding: application to hippocampus labeling. Neuroimage, 2013, 76(1): 11-23 http://europepmc.org/abstract/med/23523774 [22] Bai W J, Shi W Z, Ledig C, Rueckert D. Multi-atlas segmentation with augmented features for cardiac MR images. Medical Image Analysis, 2015, 19(1): 98-109 doi: 10.1016/j.media.2014.09.005 [23] Vapnik V N. The nature of statistical learning theory. Technometrics, 1997, 38(4): 409-409 http://portal.acm.org/citation.cfm?id=211359 [24] Smola A J. Kernel principal component analysis. In: Processings of the 1997 International Conference on Artificial Neural Networks. Springer-Verlag. 1997: 583-588 [25] Vincent P, Bengio Y. Kernel matching pursuit. Machine Learning, 2002, 48(1-3): 165-187 [26] Guigue V, Rakotomamonjy A, Canu S. Kernel basis pursuit. In: Processings of the 2005 European Conference on Machine Learning. Berlin, Heidelberg, Germany: Springer, 2005. 146-157 [27] Nguyen H V, Patel V M, Nasrabadi N M, Chellappa R. Kernel dictionary learning. In: Processings of the 2012 International Conference on Acoustics, Speech, and Signal Processing. Kyoto, Japan: IEEE, 2012. 2021-2024 [28] Drineas P, Kannan R, Mahoney M W. Fast Monte Carlo algorithms for matrices I: approximating matrix multiplication. SIAM Journal on Computing, 2006, 36(1): 132-157 doi: 10.1137/S0097539704442684 [29] Lee J, Kim S J, Chen R, Herskovits E H. Brain tumor image segmentation using kernel dictionary learning. In: Processings of the 37th Annual International Conference of the IEEE Engineering in Medicine and Biology Society. Milan, Italy: IEEE, 2015. 658-661 [30] Williams C K I, Seeger M. Using the Nyström method to speed up kernel machines. In: Processings of the 2001 Advances in Neural Information Processing Systems. 2001. 682-688 [31] Schnabel J A, Rueckert D, Quist M, Blackall J M, Castellano-Smith A D, Hartkens T. Design of non-linear discriminative dictionaries for image classification. In: Processings of the 2001 International Conference on Medical Image Computing and Computer-Assisted Intervention. Berlin, Heidelberg, Germany: Springer, 2001. 573-581 [32] Ashburner J, Friston K J. Nonlinear spatial normalization using basis functions. Human Brain Mapping, 2015, 7(4): 254-266 http://brain.oxfordjournals.org/lookup/external-ref?access_num=10408769&link_type=MED&atom=%2Fbrain%2F129%2F4%2F963.atom [33] Wang H Z, Yushkevich P A. Dependency prior for multi-atlas label fusion. In: Processings of the 9th IEEE International Symposium on Biomedical Imaging. Barcelona, Spain: IEEE, 2012. 892-895 [34] Mairal J, Bach F, Ponce J. Online dictionary learning for sparse coding. In: Processings of the 2009 International Conference on Machine Learning. Montreal, Canada: IEEE, 2009. 689-696 [35] Ojala T, Pietikäinen M, Mäenpää T. Gray scale and rotation invariant texture classification with local binary patterns. IEEE Transactions on Pattern Analysis and Machine Intelligence, 2002, 24(7): 971-987 doi: 10.1109/TPAMI.2002.1017623 [36] Ghorbani M, Targhi A T, Dehshibi M M. HOG and LBP: towards a robust face recognition system. In: Processings of the 10th International Conference on Digital Information Management. Jeju, Korea: IEEE, 2016. 138-141 [37] Zhang K, Tsang I W, Kwok J T. Improved Nyström low-rank approximation and error analysis. In: Processings of the 25th International Conference on Machine Learning. Helsinki, Finland: ACM, 2008. 1232-1239 [38] Feldman D, Feigin M, Sochen N. Learning big (image) data via coresets for dictionaries. Journal of Mathematical Imaging and Vision, 2013, 46(3): 276-291 doi: 10.1007/s10851-013-0431-x [39] Drineas P, Mahoney M W. On the Nyström method for approximating a gram matrix for improved kernel-based learning. Journal of Machine Learning Research, 2005, 6(12): 2153-2175 http://dl.acm.org/citation.cfm?id=1194916 -





 下载:
下载: